SemiCOL Challenge Submission
Challenge Tasks
The SemiCOL challenge involves two different but related tasks in digital pathology:
Task 1. The pixel-by-pixel semantic segmentation of tissue into different classes.
Task 2. The classification of whole slide images of colonic biopsies according to whether they contain tumor tissue or not (slide-level classification).
For an overview of the Challenge tasks and metrics, see Figure 1.
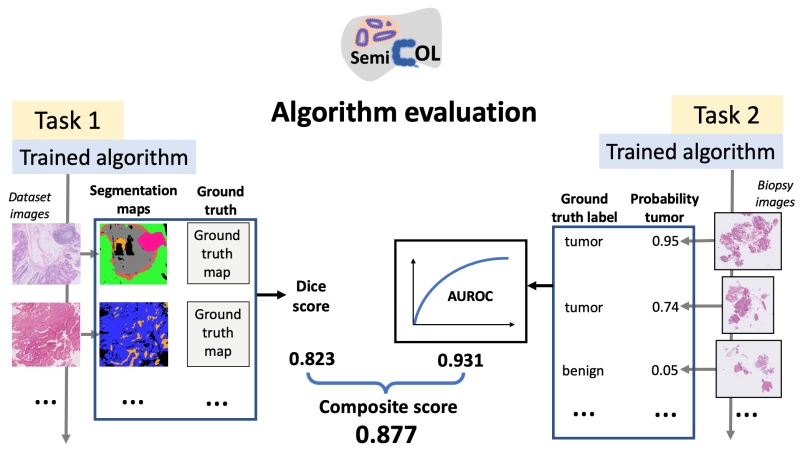
Figure 1. Overview of the SemiCOL Challenge tasks and principles of algorithm evaluation during the validation and test phase. A composite score is an average of the Dice Score and AUROC score, which are equally weighted. During Dice score calculation, all tissue classes (n=9) are weighted equally (weight = 0.1) with the exception of the tumor tissue class (weight = 0.2).
Task 1: Semantic segmentation
This task includes the generation of semantic segmentation predictions on the validation/test images for selected classes (all tissue classes, codes “1”-“9”; slide background class, BACK, code “10”, will be excluded from the generation of evaluation metrics). The metric for evaluation will be the Dice score. This will be calculated against the ground truth information (manual annotations by experienced pathologists). The evaluation metric used is the Class-weighted Dice Score. The tumor class has a class weighting of 0.2 and the rest of 0.1. The class weights for the segmentation task reflect the clinical importance of each class.
Task 2: Slide-level classification of biopsy images
For the slide-level classification task (Task 2), the area under the receiver operating curve (AUROC) for slide-level probability to contain a tumor provided by participants' algorithms is used. Therefore, the output of the algorithm should include the probability of the whole slide containing a tumor or not.
Task 1 and 2: Composite metric for algorithm ranking
As there are multiple test datasets involved in both Task 1 and Task 2, Dice Score and AUROC metrics for different datasets will be averaged resulting in mean Dice Score and mean AUROC. Then a final score will be computed by averaging these mean Dice Score and AUROC. The ranking will be done based on that averaged value. An equal weighting for both metrics has been chosen, as we consider both tasks to be as important as one another.
Submission during Validation phase
Required data structure
Since the SemiCOL challenge is about both segmentation and tumor classification, two different types of predictions must be submitted:
The predicted segmentation maps for the images from the manually annotated dataset (/01_MANUAL/wns_case_01..06). Submission: Segmentation maps.
The predicted tumor probability for the 40 biopsy slides (/02_BX/...). Submission: JSON file with slide-level probabilities.
The compressed (.zip) submission folder should be uploaded into the teams Google drive "Validation" folder. The folder name will also be used to name the response file. The folder should have the following structure:
|-- classification.json
|-- wns_case_01
| |-- wns_case_01 [d=2.16945,x=45558,y=26033,w=6509,h=6509].png
| |-- wns_case_01 [d=2.16945,x=45558,y=32542,w=6509,h=6508].png
| |-- ...
|-- wns_case_02
| |-- wns_case_02A [d=2.16945,x=26033,y=39050,w=6509,h=6508].png
| |-- ...
|-- ...
Segmentation maps
- The maps must be saved as .png files
- The maps must be readable with pillow Image.open() and should result in a matrix with size (h x w)
- The maps must have the same size as the images. Otherwise, the map will be excluded.
- Missing or excluded predictions will be set to a false prediction for all pixels
- The map has to be named like the corresponding image (files with different names will be ignored)
- The folder structure must be the same as for the validation images
- The maps should only contain values between 1 and 10.
Classification JSON file
To calculate the AUC for the classification task, a json file containing the predicted probabilities for single biopsy slides (ome.tiff files) must be included. It must have the following structure:
{
"ABLVABMT": 0.9,
"QgiI3ZjZ": 0.3,
"4Hjkjs0i": 0.1,
...
}
- The JSON file must be named classification.json
- Missing slides will receive a false prediction (0 if the label is 1 and 1 if the label is 0)
- All probabilities will be clipped to the range [0,1]
- Including any other file names will be ignored
Response
The response will be uploaded into your Google drive "Validation" folder and will contain the team key, arm and the averaged score. It will also contain information about the individual tasks. If probabilities are missing or if maps contain invalid numbers, this will also be listed in the task specific sections.
{
"private_key": "12345678",
"arm": 1,
"score": 0.734375,
"classification": {
"auc": 0.46875,
"missing": ["BHTUesDo", "x2cYOIty"]
},
"segmentation": {
"dice": 1.0,
"missing": [],
"irregular_patches": [
[
"01_WNS",
"wns_case_01",
"wns_case_01 [d=2.16945,x=45558,y=26033,w=6509,h=6509].png"
]
]
}
}
In case of an error, the response will have the following form:
{"error": "Error Message"}
Docker instructions
Runtime requirements
During the test phase, we will run your docker images on a PC with an i7-13700K CPU, 128 GB RAM and a 4090RTX GPU. The computer will be offline and will not have any connection to the internet.
Running your docker image
When we receive your docker tar file, we will load and run your algorithm with the following commands:
docker load < teamname.tar.gz
docker run --gpus "device=0" –name teamname –v /home/semicol/input:/input –v /home/semicol/predict:/predict teamname:latest
This means that we will mount the folder /home/semicol/input (containing all input files for testing) to /input in your docker container, and mount the folder /home/semicol/predict (an empty folder used to save your predictions) to /predict in your docker container.
The data structure in /input will be as follows:
├── input
│ ├── 01_MANUAL
│ │ ├── [case folders]
│ │ │ ├── *.png
│ ├── 02_BX
│ │ ├── *.ome.tiff
In summary, your algorithm should:
- Parse the content of
/inputto find the images that to be either segmented or classified. - Run inference.
- Save the predictions in
/predict. Note that the overall structure of the predictions and the naming conventions are the same as during the validation phase.
Saving your docker image
Please name your docker image as your team name, and set the tag to "latest". Then save your docker image as teamname.tar.gz. For instance, you can use this command:
docker save teamname:latest –o teamname.tar.gz
Note: docker image name has to be in lowercase.
Submitting your docker image
To submit your docker image, simply upload it to the root directory of your team Google Drive folder. This is the folder structure where you can submit predictions during the validation phase.
Submission during the Test phase
All test datasets will remain private. The process of evaluation will be carried out on the Organizers’ side. The participating teams should provide the following data during submission:
1. Containerized version of the algorithm. Includes information on Challenge Arm.
2. Detailed description of the algorithm (pdf), of the data/data preprocessing used.
3. Raw code of the algorithm.
4. Any additional annotations used (Arm 2).
The guidelines for algorithm submission will be provided on the March 1st 2023.
Technical issues with submission evaluation
If a submission is missing or the algorithm does not work as intended, the submission will be excluded from any ranking procedures. If the algorithm does not work for a given slide / slides, then a Dice-Score of 0 will be assumed and the wrong classification label with confidence of 1 will be used for the AUROC calculation.
Ranking of the algorithms
Ranking of the algorithms will be based on the Challenge Arm (Arm 1 or 2) and the Composite metric (see above). The winners will be determined in each Arm (Arm 1 and 2). Should one team be among the winners in both Arms, it can be nominated as the winner only in one Arm (the highest place). It will be excluded from nomination in the other Arm, leading to upstaging of the teams with the next following top-performing algorithms.
Additional evaluation of the winning algorithms
The organizers retain a right to perform a complete retraining/evaluation of the top performing/winning algorithms on their own infrastructure (Arms 1, 2). The denial to provide all necessary data: raw code, data, data preprocessing pipeline, validation pipeline, seeds, additional annotations, run instructions, or any other essential components (checkpoints/weights, etc.) will result in the exclusion of the algorithm from winner nominations.